About the Department of Statistics
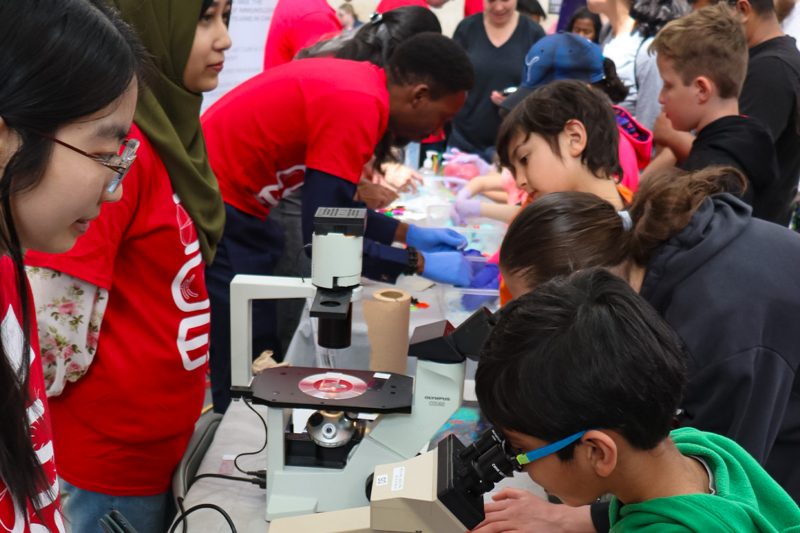
Statistics lives at the core of scientific reasoning, discovery and development, and its expertise can be used to shape public policy and business decisions.
The Department of Statistics is one of the oldest and largest in Canada. Most of our upper-level statistical courses are accredited by the Statistical Society of Canada toward the Associate Statistician (AStat) designation. We have a high academic staff-to-student ratio, giving our students greater personal attention and more opportunities to tailor their educational experiences.
The department is home to an immersive research environment, and in collaboration with the Canadian Statistical Sciences Institute (CANSSI) has established a Regional Research Centre for the Prairie Region: Manitoba, Saskatchewan, NWT, and Nunavut (CANSSI Prairies) at UM to further promote statistical sciences research and training, both in Canada and internationally.