Antimicrobial resistance
- Global priority
- National surveillance
- Resistance mechanisms
The University of Manitoba campuses are located on original lands of Anishinaabeg, Ininew, Anisininew, Dakota and Dene peoples, and on the National Homeland of the Red River Métis. More
University of Manitoba
Winnipeg, Manitoba Canada, R3T 2N2
The Department of Medical Microbiology and Infectious Diseases is an integrated team of investigators, educators and practitioners providing leading edge education, innovative research and expertise in medical microbiology and infectious diseases.
We are seen nationally and internationally as dynamic leaders and collaborators. We are acknowledged for excellence in numerous areas and are constantly developing new research initiatives in infectious diseases.
Our efforts create and translate new knowledge, shaping policy and contributing to economic growth to improve the lives of those in the region and throughout the world.
The department supports a number of education programs, from undergraduate studies to post-doctoral training in specialty areas of microbiology.
Established in 1884 as the Department of Bacteriology, Pathology, and Histology, our department has a rich and evolving history.
It underwent several transformations, becoming Bacteriology and Hygiene in 1916, Bacteriology and Serology in 1919, Bacteriology and Immunology in 1929, and finally adopting the name Medical Microbiology in 1967. In 2016, the department evolved into Medical Microbiology and Infectious Diseases (MMID), reflecting a heightened focus on both the pathogen and the patient's disease.
With a legacy dating back over a century, MMID remains at the forefront of medical research and care. The department seamlessly integrates robust basic science research and graduate student training with a commitment to excellence in clinical microbiology, infectious disease care, and postgraduate clinical training.
Watch a brief video to learn more about our department and what we offer.

UM is a community that cares. We offer a wide variety of supports to help you make the most of your time at university and reach your full potential.
The Manitoba Medical College Foundation (MMCF) has established the MMCF – Jack Wilt Travel Award at the Winnipeg Foundation through gifts from Dr. John Charles (Jack) Wilt's friends, family, medical institutions, and conferences. This award aims to assist students with travel expenses for conference participation, funded by the annual income generated from the endowment.
To qualify for the award, applicants must be in good standing and attending a professional meeting or conference to present research results (poster or oral presentation). Eligible candidates are enrolled in either the postgraduate medical education program in the Department of Medical Microbiology at the Max Rady College of Medicine or the Adult or Pediatric Infectious Disease Residency Training Program at the same college. Requests for funding should include evidence of paper or poster presentation acceptance.
The dean of the Max Rady College of Medicine (or designate) will appoint the selection committee, including the head of medical microbiology and infectious diseases (or designate). The University of Manitoba's board of governors retains the right to modify award terms with complete consultation with the Winnipeg Foundation if changed conditions necessitate adjustments.
Financial aid and awards send the award selection form to the PGME awards coordinator.
The PGME awards coordinator forwards relevant award details to the department-based award administrator.
After selecting a candidate, the department-based awards administrator provides the resident's name, student number, and award amount to the PGME awards coordinator.
The PGME awards coordinator sends a signed, completed award selection form back to the Financial Aid and Awards Office for processing.
Award status information cannot be provided by the department or PGME office via email, phone, or in person.
The Financial Aid and Awards Office communicates the award to the recipient through a notification letter. Once the cheque is issued, it goes to the PGME awards coordinator for disbursement to the department-based award administrator, who then forwards it to the successful candidate.
The department produces excellent research to continually develop our understanding of microbiology and infectious diseases. Some of our research strengths include:
Explore the links below to learn more about our groundbreaking researchers, including:
We are proud to partner with organizations worldwide to improve public health in the field of infectious diseases on an international scale.
Our department is run by award-winning faculty and staff who are dedicated and passionate about their contributions to the field of medical microbiology. Contact us to learn more about our department and what we have to offer.
Position: Assistant professor – probationary (tenure track)
Deadline to apply: November 15, 2023
Start date: March 1, 2024
The Department of Medical Microbiology and Infectious Diseases (MMID) invites applications for a full-time tenure-track Assistant Professor in Emerging Pathogens, commencing March 1, 2024 or on a date mutually agreed upon. Salary will be commensurate with experience and qualifications.
The Department seeks an emerging scholar with a commitment to excellence in teaching and research. Exceptional candidates will be considered with the responsibilities and qualifications outlined below:
Description and responsibilities:
The duties of this position include teaching, research, and service.
Teaching
Teaching undergraduate and graduate students and mentoring/supervising research trainees.
Research
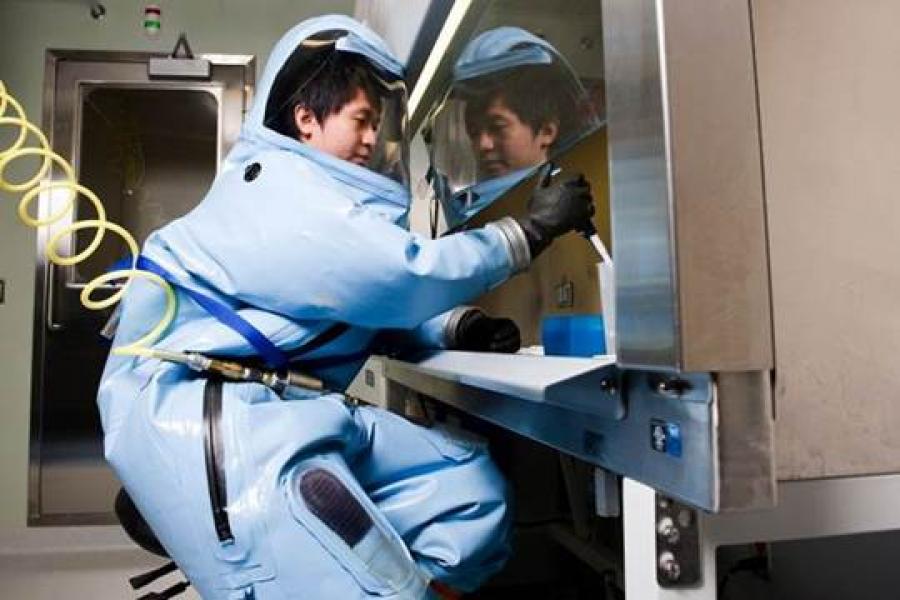
Develop and maintain a rigorous, independent, nationally funded, research program that is highly collaborative involving local, national, and international partners. The program should be highly collaborative with members of MMID, the University and regional partners such as local hospitals, the provincial laboratory and the National Microbiology Laboratory. The focus of this position is on containment level 3 emerging pathogens. The program must involve in vitro CL3 work utilizing the new in vitro laboratory built by MMID for this purpose. The possibility of animal CL3 work will be considered but must involve well defined relationships with external partners as the UM currently does not have animal CL3 capability.
Service
Service to the department, Max Rady College of Medicine, University, outreach activities including service to the external scientific community through manuscript and grant reviews.
Qualifications
The successful candidate must have a Ph.D. and/or MD with postdoctoral research experience and a productive track record of peer-reviewed publications. Experience with high containment pathogens is required.
Teaching and student mentorship experience preferred.
The successful candidate’s research program must involve emerging pathogens and must align with one or more of the department's identified areas of research strength (HIV, viral pathogenesis, host/pathogen interactions and antimicrobial resistance).
About Us
The Department of MMID currently has ten full-time tenured and tenure-track faculty members and 76 affiliated members and offers MSc and PhD graduate programs in Medical Microbiology and Infectious Diseases with a current enrollment of over 60 students.
The University of Manitoba is committed to the principles of equity, diversity & inclusion and to promoting opportunities in hiring, promotion and tenure (where applicable) for systemically marginalized groups who have been excluded from full participation at the University and the larger community including Indigenous Peoples, women, racialized persons, persons with disabilities and those who identify as 2SLGBTQIA+ (Two Spirit, lesbian, gay, bisexual, trans, questioning, intersex, asexual and other diverse sexual identities). All qualified candidates are encouraged to apply; however, Canadian citizens and permanent residents will be given priority.
An inclusive, open and diverse community is essential to excellence and fosters voices that have been ignored or discouraged. To address the Rady Faculty of Health Sciences commitment to equity, diversity and inclusion, and in recognition of the underrepresentation of members of historically and currently excluded groups, we take proactive measures including implicit bias training for all hiring panels. We strive for diversity and cultural safety throughout the hiring process (hiring panels, short-list of candidates, interviews). We encourage you to self-identify any aspect of diversity in your cover letter.
If you require accommodation supports during the recruitment process, please contact UM.Accommodation@umanitoba.ca or 204-474-7195. Please note this contact information is for accommodation reasons only.
Application materials must include:
Send to:
Dr. Keith Fowke
Search Committee Chair
544-745 Bannatyne Ave Winnipeg, MB, R3E 0J9
c/o Leah Buermeyer
leah.buermeyer@umanitoba.ca
The closing date for receipt of applications is November 15, 2023.
Application materials, including letters of reference, will be handled in accordance with the protection of privacy provision of The Freedom of Information and Protection of Privacy Act (Manitoba). Please note that curriculum vitae may be provided to participating members of the search process.

Faculty of Agricultural and Food Sciences, UM Today

Rady Faculty of Health Sciences, Research and International, Students

Rady Faculty of Health Sciences, UM Today
Medical Microbiology and Infectious Diseases
Max Rady College of Medicine
Room 543 - 745 Bannatyne Avenue
University of Manitoba (Bannatyne campus)
Winnipeg, MB R3E 0J9 Canada